One-to-n alignments
Command: compare-matrices -v 1 -mode matches -format1 transfac -file1 $RSAT/public_html/tmp/wwwrun/2013/01/11/peak-motifs.2013-01-11.110705_2013-01-11.110705_PdhmNx/results/discovered_motifs/oligos_7nt_mkv4_m1/peak-motifs_oligos_7nt_mkv4_m1.tf -format2 tf -file2 $RSAT/public_html/data/motif_databases/JASPAR/jaspar_core_vertebrates_2009_10.tf -mode matches -DR -uth offset_rank 1 -lth w 5 -lth Wr 0.3 -lth cor 0.75 -lth Ncor 0.4 -return matrix_name,matrix_id,cor,Ncor,logoDP,NIcor,NsEucl,SSD,NSW,match_rank,width,strand,offset,consensus,alignments_1ton -sort Ncor -o $RSAT/public_html/tmp/wwwrun/2013/01/11/peak-motifs.2013-01-11.110705_2013-01-11.110705_PdhmNx/results/discovered_motifs/oligos_7nt_mkv4_m1/peak-motifs_oligos_7nt_mkv4_m1_vs_db_jaspar_core_vertebrates
One-to-n matrix alignment; reference matrix: oligos_7nt_mkv4_m1_shift0 ; 7 matrices ; sort_field=rank_mean
| Matrix name | Aligned logos | cor |
Ncor |
logoDP |
NIcor |
NsEucl |
SSD |
NSW |
rcor |
rNcor |
rlogoDP |
rNIcor |
rNsEucl |
rSSD |
rNSW |
rank_mean |
match_rank |
Aligned matrices |
|---|
| oligos_7nt_mkv4_m1_shift0 (oligos_7nt_mkv4_m1) |
 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
; oligos_7nt_mkv4_m1; m=0 (reference); ncol1=14; shift=0; ncol=15; srTTTCCwGGAAay-
; Alignment reference
a 237 426 84 1 1 178 0 456 265 0 1319 1318 550 257 0
c 455 268 57 1 40 1142 1320 266 1 0 0 2 202 358 0
g 349 370 74 0 0 0 0 176 1054 1138 0 0 315 228 0
t 279 256 1105 1318 1279 0 0 422 0 182 1 0 253 477 0
|
| MA0137.2_shift0 (STAT1) |
 |
0.967 |
0.903 |
9.138 |
0.901 |
0.969 |
0.372 |
0.987 |
1 |
1 |
1 |
1 |
1 |
1 |
1 |
1.000 |
1 |
; oligos_7nt_mkv4_m1 versus MA0137.2 (STAT1); m=1/6; ncol2=15; w=14; offset=0; strand=D; shift=0; score= 1; cayTTCChrGAAryc
; cor=0.967; Ncor=0.903; logoDP=9.138; NIcor=0.901; NsEucl=0.969; SSD=0.372; NSW=0.987; rcor=1; rNcor=1; rlogoDP=1; rNIcor=1; rNsEucl=1; rSSD=1; rNSW=1; rank_mean=1.000; match_rank=1
a 208 859 251 10 8 106 23 528 696 53 1900 2030 954 336 417
c 1076 496 574 22 14 1921 1900 762 31 30 124 17 263 760 804
g 415 279 144 11 38 14 7 115 1292 1700 29 23 552 270 425
t 378 446 1112 2038 2023 44 155 680 66 302 32 15 315 714 431
|
| MA0144.1_shift3 (Stat3) |
 |
0.948 |
0.677 |
8.999 |
0.673 |
0.948 |
0.541 |
0.973 |
2 |
2 |
2 |
2 |
2 |
2 |
2 |
2.000 |
2 |
; oligos_7nt_mkv4_m1 versus MA0144.1 (Stat3); m=2/6; ncol2=10; w=10; offset=3; strand=D; shift=3; score= 2; ---TTCCaGGAAr--
; cor=0.948; Ncor=0.677; logoDP=8.999; NIcor=0.673; NsEucl=0.948; SSD=0.541; NSW=0.973; rcor=2; rNcor=2; rlogoDP=2; rNIcor=2; rNsEucl=2; rSSD=2; rNSW=2; rank_mean=2.000; match_rank=2
a 0 0 0 20 13 38 6 321 8 6 585 606 191 0 0
c 0 0 0 19 10 552 541 21 0 2 21 1 15 0 0
g 0 0 0 25 129 9 1 148 605 592 7 5 393 0 0
t 0 0 0 549 461 14 65 123 0 13 0 1 14 0 0
|
| MA0152.1_shift1 (NFATC2) |
 |
0.913 |
0.456 |
6.329 |
0.454 |
0.919 |
0.639 |
0.954 |
3 |
5 |
3 |
5 |
3 |
3 |
3 |
3.571 |
3 |
; oligos_7nt_mkv4_m1 versus MA0152.1 (NFATC2); m=3/6; ncol2=7; w=7; offset=1; strand=D; shift=1; score= 3.5714; -TTTTCCA-------
; cor=0.913; Ncor=0.456; logoDP=6.329; NIcor=0.454; NsEucl=0.919; SSD=0.639; NSW=0.954; rcor=3; rNcor=5; rlogoDP=3; rNIcor=5; rNsEucl=3; rSSD=3; rNSW=3; rank_mean=3.571; match_rank=3
a 0 3 1 1 0 1 0 18 0 0 0 0 0 0 0
c 0 1 2 1 0 25 26 3 0 0 0 0 0 0 0
g 0 2 2 0 0 0 0 1 0 0 0 0 0 0 0
t 0 20 21 24 26 0 0 4 0 0 0 0 0 0 0
|
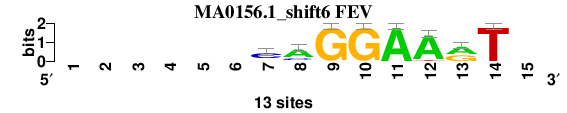
| MA0156.1_shift6 (FEV) |
 |
0.858 |
0.491 |
5.956 |
0.468 |
0.905 |
1.161 |
0.927 |
4 |
3 |
4 |
4 |
4 |
5 |
5 |
4.143 |
4 |
; oligos_7nt_mkv4_m1 versus MA0156.1 (FEV); m=4/6; ncol2=8; w=8; offset=6; strand=D; shift=6; score= 4.1429; ------CAGGAArT-
; cor=0.858; Ncor=0.491; logoDP=5.956; NIcor=0.468; NsEucl=0.905; SSD=1.161; NSW=0.927; rcor=4; rNcor=3; rlogoDP=4; rNIcor=4; rNsEucl=4; rSSD=5; rNSW=5; rank_mean=4.143; match_rank=4
a 0 0 0 0 0 0 2 9 0 0 13 12 7 0 0
c 0 0 0 0 0 0 9 3 0 0 0 0 0 0 0
g 0 0 0 0 0 0 1 1 13 13 0 0 6 0 0
t 0 0 0 0 0 0 1 0 0 0 0 1 0 13 0
|
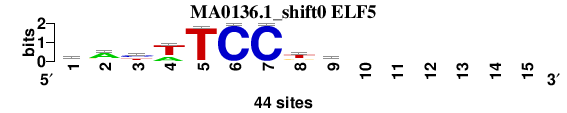
| MA0136.1_shift0 (ELF5) |
 |
0.760 |
0.488 |
5.328 |
0.495 |
0.899 |
1.651 |
0.908 |
6 |
4 |
5 |
3 |
5 |
6 |
6 |
5.000 |
5 |
; oligos_7nt_mkv4_m1 versus MA0136.1 (ELF5); m=5/6; ncol2=9; w=9; offset=0; strand=D; shift=0; score= 5; waywTCCkk------
; cor=0.760; Ncor=0.488; logoDP=5.328; NIcor=0.495; NsEucl=0.899; SSD=1.651; NSW=0.908; rcor=6; rNcor=4; rlogoDP=5; rNIcor=3; rNsEucl=5; rSSD=6; rNSW=6; rank_mean=5.000; match_rank=5
a 12 29 2 14 0 0 0 4 4 0 0 0 0 0 0
c 9 3 20 0 1 44 44 3 8 0 0 0 0 0 0
g 3 7 5 1 0 0 0 11 11 0 0 0 0 0 0
t 20 5 17 29 43 0 0 26 21 0 0 0 0 0 0
|
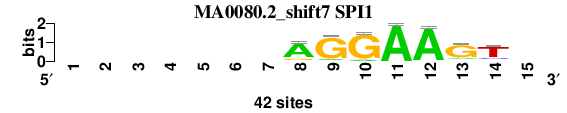
| MA0080.2_shift7 (SPI1) |
 |
0.855 |
0.427 |
5.088 |
0.423 |
0.899 |
1.000 |
0.929 |
5 |
6 |
6 |
6 |
6 |
4 |
4 |
5.286 |
6 |
; oligos_7nt_mkv4_m1 versus MA0080.2 (SPI1); m=6/6; ncol2=7; w=7; offset=7; strand=D; shift=7; score= 5.2857; -------AGGAAGT-
; cor=0.855; Ncor=0.427; logoDP=5.088; NIcor=0.423; NsEucl=0.899; SSD=1.000; NSW=0.929; rcor=5; rNcor=6; rlogoDP=6; rNIcor=6; rNsEucl=6; rSSD=4; rNSW=4; rank_mean=5.286; match_rank=6
a 0 0 0 0 0 0 0 33 4 4 42 41 5 3 0
c 0 0 0 0 0 0 0 3 0 0 0 0 3 5 0
g 0 0 0 0 0 0 0 6 37 38 0 1 33 2 0
t 0 0 0 0 0 0 0 0 1 0 0 0 1 32 0
|






