One-to-n alignments
Command: compare-matrices -v 1 -mode matches -format1 transfac -file1 $RSAT/public_html/tmp/wwwrun/2013/01/11/peak-motifs.2013-01-11.110705_2013-01-11.110705_PdhmNx/results/discovered_motifs/positions_7nt_m1/peak-motifs_positions_7nt_m1.tf -format2 tf -file2 $RSAT/public_html/data/motif_databases/JASPAR/jaspar_core_vertebrates_2009_10.tf -mode matches -DR -uth offset_rank 1 -lth w 5 -lth Wr 0.3 -lth cor 0.75 -lth Ncor 0.4 -return matrix_name,matrix_id,cor,Ncor,logoDP,NIcor,NsEucl,SSD,NSW,match_rank,width,strand,offset,consensus,alignments_1ton -sort Ncor -o $RSAT/public_html/tmp/wwwrun/2013/01/11/peak-motifs.2013-01-11.110705_2013-01-11.110705_PdhmNx/results/discovered_motifs/positions_7nt_m1/peak-motifs_positions_7nt_m1_vs_db_jaspar_core_vertebrates
One-to-n matrix alignment; reference matrix: positions_7nt_m1_shift0 ; 4 matrices ; sort_field=rank_mean
| Matrix name | Aligned logos | cor |
Ncor |
logoDP |
NIcor |
NsEucl |
SSD |
NSW |
rcor |
rNcor |
rlogoDP |
rNIcor |
rNsEucl |
rSSD |
rNSW |
rank_mean |
match_rank |
Aligned matrices |
|---|
| positions_7nt_m1_shift0 (positions_7nt_m1) |
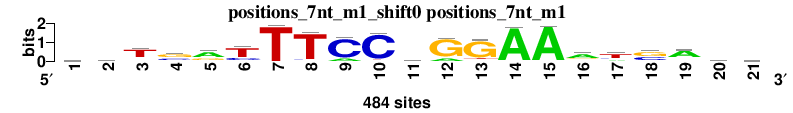 |
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
|
; positions_7nt_m1; m=0 (reference); ncol1=21; shift=0; ncol=21; wsTsrTTTCCwGGAAatsAsa
; Alignment reference
a 124 107 52 35 289 18 1 4 77 11 153 80 33 472 477 287 85 17 330 114 138
c 117 138 39 167 31 113 2 28 401 442 92 2 4 8 3 30 87 196 57 143 117
g 112 148 53 247 137 19 2 6 2 3 97 395 400 2 2 105 23 244 51 131 113
t 131 91 340 35 27 334 479 446 4 28 142 7 47 2 2 62 289 27 46 96 116
|
| MA0137.2_rc_shift3 (STAT1_rc) |
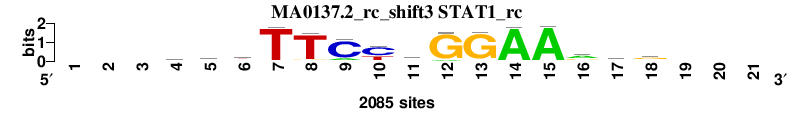 |
0.945 |
0.675 |
7.824 |
0.676 |
0.963 |
0.625 |
0.979 |
2 |
1 |
1 |
1 |
1 |
2 |
1 |
1.286 |
1 |
; positions_7nt_m1 versus MA0137.2_rc (STAT1_rc); m=1/3; ncol2=15; w=15; offset=3; strand=R; shift=3; score= 1.2857; ---gryTTCydGGAArtg---
; cor=0.945; Ncor=0.675; logoDP=7.824; NIcor=0.676; NsEucl=0.963; SSD=0.625; NSW=0.979; rcor=2; rNcor=1; rlogoDP=1; rNIcor=1; rNsEucl=1; rSSD=2; rNSW=1; rank_mean=1.286; match_rank=1
a 0 0 0 431 714 315 15 32 302 66 680 155 44 2023 2038 1112 446 378 0 0 0
c 0 0 0 425 270 552 23 29 1700 1292 115 7 14 38 11 144 279 415 0 0 0
g 0 0 0 804 760 263 17 124 30 31 762 1900 1921 14 22 574 496 1076 0 0 0
t 0 0 0 417 336 954 2030 1900 53 696 528 23 106 8 10 251 859 208 0 0 0
|
| MA0144.1_shift6 (Stat3) |
 |
0.948 |
0.451 |
7.793 |
0.447 |
0.948 |
0.531 |
0.973 |
1 |
2 |
2 |
2 |
2 |
1 |
2 |
1.714 |
2 |
; positions_7nt_m1 versus MA0144.1 (Stat3); m=2/3; ncol2=10; w=10; offset=6; strand=D; shift=6; score= 1.7143; ------TTCCaGGAAr-----
; cor=0.948; Ncor=0.451; logoDP=7.793; NIcor=0.447; NsEucl=0.948; SSD=0.531; NSW=0.973; rcor=1; rNcor=2; rlogoDP=2; rNIcor=2; rNsEucl=2; rSSD=1; rNSW=2; rank_mean=1.714; match_rank=2
a 0 0 0 0 0 0 20 13 38 6 321 8 6 585 606 191 0 0 0 0 0
c 0 0 0 0 0 0 19 10 552 541 21 0 2 21 1 15 0 0 0 0 0
g 0 0 0 0 0 0 25 129 9 1 148 605 592 7 5 393 0 0 0 0 0
t 0 0 0 0 0 0 549 461 14 65 123 0 13 0 1 14 0 0 0 0 0
|
| MA0062.2_shift9 (GABPA) |
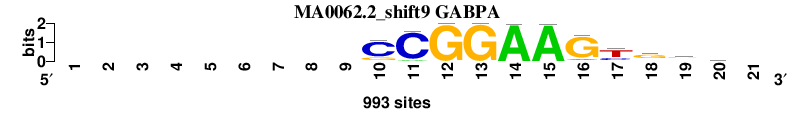 |
0.778 |
0.408 |
5.981 |
0.399 |
0.909 |
2.004 |
0.909 |
3 |
3 |
3 |
3 |
3 |
3 |
3 |
3.000 |
3 |
; positions_7nt_m1 versus MA0062.2 (GABPA); m=3/3; ncol2=11; w=11; offset=9; strand=D; shift=9; score= 3; ---------CCGGAAGygvc-
; cor=0.778; Ncor=0.408; logoDP=5.981; NIcor=0.399; NsEucl=0.909; SSD=2.004; NSW=0.909; rcor=3; rNcor=3; rlogoDP=3; rNIcor=3; rNsEucl=3; rSSD=3; rNSW=3; rank_mean=3.000; match_rank=3
a 0 0 0 0 0 0 0 0 0 32 70 0 0 991 989 94 56 154 264 233 0
c 0 0 0 0 0 0 0 0 0 768 914 0 0 1 2 32 261 137 262 357 0
g 0 0 0 0 0 0 0 0 0 188 4 990 991 1 0 866 37 603 415 224 0
t 0 0 0 0 0 0 0 0 0 1 1 1 0 0 2 0 637 96 49 176 0
|



